 Convergence of genes and cellular pathways dysregulated in autism spectrum disorders. Functional impact of global rare copy number variation in autism spectrum disorders. Epigenomic alterations define lethal CIMP-positive ependymomas of infancy. Big data: astronomical or genomical? PLoS Biol. Initial impact of the sequencing of the human genome. The complete protocol can be performed in ~4.5 h and is designed for use by biologists with no prior bioinformatics training. The protocol describes innovative visualization techniques, provides comprehensive background and troubleshooting guidelines, and uses freely available and frequently updated software, including g:Profiler, Gene Set Enrichment Analysis (GSEA), Cytoscape and EnrichmentMap. We describe how to use this protocol with published examples of differentially expressed genes and mutated cancer genes however, the principles can be applied to diverse types of omics data. 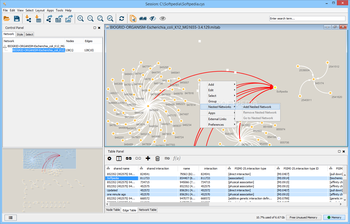
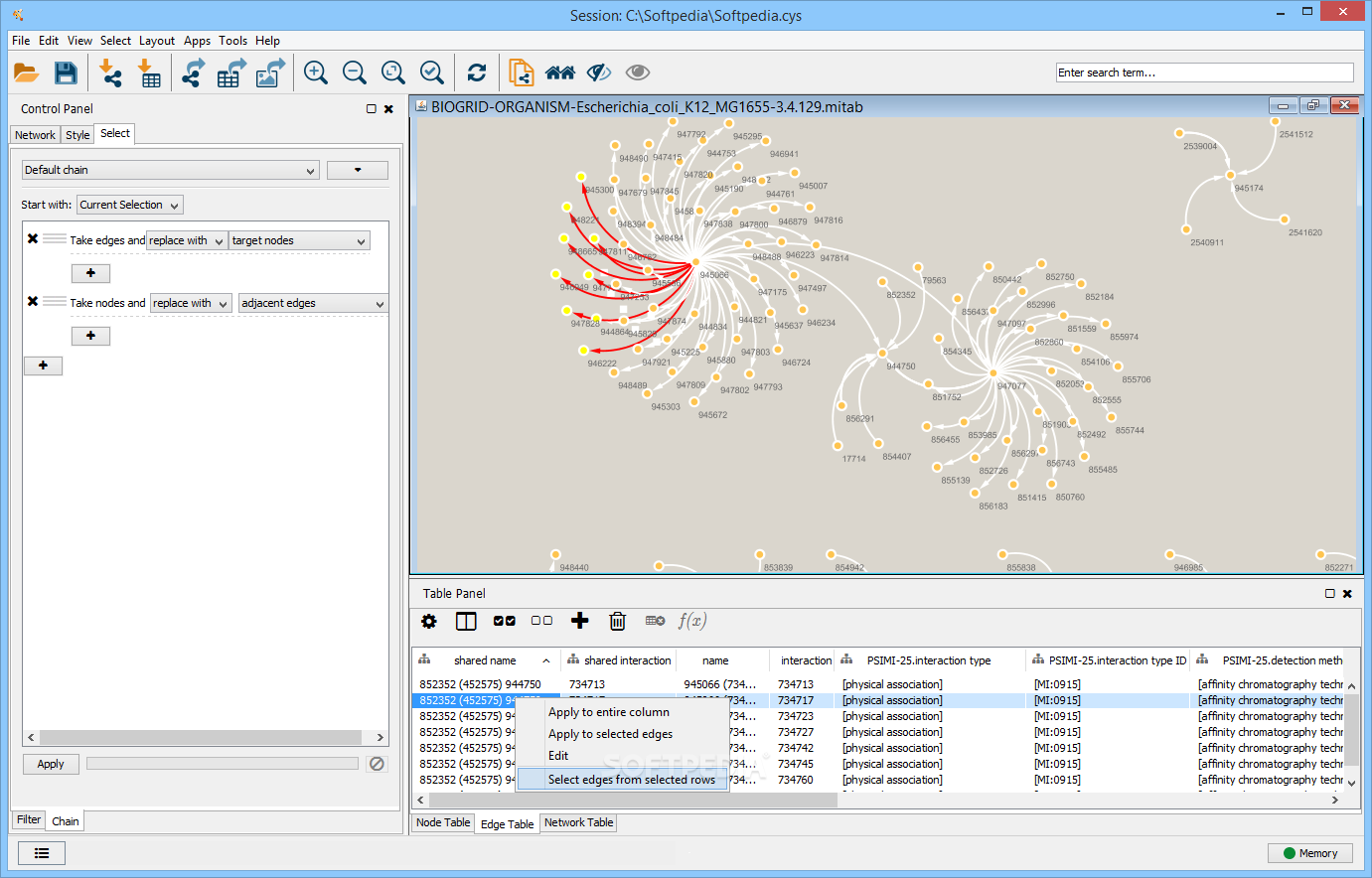
The protocol comprises three major steps: definition of a gene list from omics data, determination of statistically enriched pathways, and visualization and interpretation of the results. We explain the procedures of pathway enrichment analysis and present a practical step-by-step guide to help interpret gene lists resulting from RNA-seq and genome-sequencing experiments. This method identifies biological pathways that are enriched in a gene list more than would be expected by chance. Please note that if you are not eligible for a University of Cambridge Raven account you will need to book or register your interest by linking here.Pathway enrichment analysis helps researchers gain mechanistic insight into gene lists generated from genome-scale (omics) experiments. The course will also access and analyse the data through Cytoscape apps, including the new IntAct app. The course will focus on giving attendees hands-on experience in the use of Cytoscape, an open source software platform for complex network analysis and visualization. We will subsequently use these networks to overlay large-scale data such as that obtained through RNA-Seq or mass-spec proteomics. Attendees will learn how to construct protein-protein interaction networks and explore them in full detail. This course provides an introduction to the basic theory and concepts of network analysis. We aim to simulate the classroom experience as closely as possible, with opportunities for one-to-one discussion with tutors and a focus on interactivity throughout. PLEASE NOTE The Bioinformatics Team are presently teaching as many courses live online.
0 Comments
Leave a Reply. |

 RSS Feed
RSS Feed
